As the pioneer of liquid chip application in agriculture, Molbreeding is committed to improving the detection capacity, reducing test cost and improving the efficiency of molecular breeding of animals and plants. MGI and Molbreeding have reached a strategic cooperation to further promote molecular breeding innovation and industrialization.

Recently, Molbreeding evaluated the performance of the porcine 50K panel on MGI's DNBSEQ-T7 gene sequencer*, including the operation of the DNBSEQ-T7*, target area data coverage, uniformity, and genotype effect.

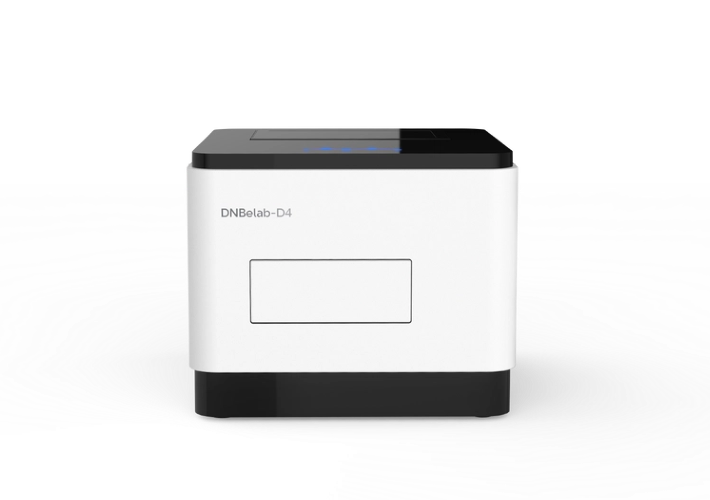
MGI Genetic Sequencers DNBSEQ-400* (left) and DNBSEQ-T7* (right)
Materials and methods
Liquid chip: PHR0101_Sus50K
Product name: GenoBaits® Porcine 50K Panel
Number of marks: 52,000
Recommended data: 2.2G/ samples
Average coverage depth of target area: 105.88x
Library Preparation Method: DNA library pre + probe-in-solution -based (liquid chip)
Sequencing platform: DNBSEQ-T7
1. High data validity
1.1 High Mapping Rate
Mapping Rate reflects the similarity between the sample sequencing data and the reference genome and is an indicator of the overall sequencing accuracy and the presence of contamination. The higher mapping rate, the better, indicating high data validity. In this evaluation, 10 samples showed excellent mapping rate on DNBSEQ-T7 gene sequencer* with the mapping rate of each sample higher than 99.50%. As shown in the figure below.

Figure1. Data Mapping Rate Based on DNBSEQ-T7 Gene Sequencer*
1.2 Excellent Capture Efficiency of Target Region
Capture Rate/Efficiency refers to the ratio of aligned sequences on target region to all sequences aligned to the reference genome. The capture efficiency does not affect the data quality, but only the effective ratio of data. The higher the capture efficiency, the smaller the amount of data required to cover the same depth, and the lower the sequencing cost, indicating high data validity. In this evaluation, the capture efficiency of 10 samples is about 62%-63%, and the highest is about 63.82%. As shown in the figure below.

Fig. 2. Data Capture Efficiency Based on DNBSEQ-T7 Gene Sequencer*
2 High Coverage and Uniformity of Target Region
Coverage (depth) refers to the percentage of sequencing data to be located on the sequence of the target region after splicing and comparison. It reflects the sequencing coverage and uniformity of the target region. With the same target region and same number of aligned sequences reads, the more uniform and higher the coverage reflects more uniform sequence coverage of the target region. Based on DNBSEQ-T7* in this evaluation, the average uniformity_rate10 of 10 samples was 99.56%, and the uniformity_rate10 of all 10 samples was higher than 99.40%, the highest being 99.69%. The average uniformity_20 of 10 samples is 96.70%, the uniformity_20 of 10 samples is higher than 96.10%, and the highest uniformity_20 of 10 samples is 97.27%.

Fig. 3. Data Uniformity Based on DNBSEQ-T7*
3 High-quality Genotyping
3.1 High SNP Detection Rate
SNP detection rate and individual detection rate are important measurements of chip quality, which are generally measured by the loci of autosomal and X chromosome. The detection rates of all 10 samples are higher than 99.71%, with the maximum detection rate of 99.92% and the minimum detection rate of 99.71%, and the average detection rate of 99.8%. All 10 samples show a relatively high detection rate.

Fig. 4. SNP Detection Rate Based on DNBSEQ-T7 *
3.2 Stable Genotype Detection
Generally, the stability is measured by the consistency and correlation coefficient of two genotype detection results of duplicate samples. In this evaluation, six test samples were selected for genotypic consistency detection, and the results of two genotypic tests of six duplicate samples were as follows: the average genotype consistency was 99.22%, and the highest genotype consistency reached 99.27%, which indicated that the product PHR0101_Sus50K_V1.0 had good stability on DNBSEQ-T7*.

Fig. 5. Genotype consistency based on DNBSEQ-T7*
4.Uniform Distribution Map of 4 PHR0101_Sus50K_V1.0 Loci
In this evaluation, SNP markers detected by liquid chip PHR0101_Sus50K_V1.0 based on DNBSEQ-T7* are evenly distributed on each chromosome.

Fig. 6. SNP markers based on DNBSEQ-T7* are evenly distributed on each chromosome.
Evaluation Summary
From the above evaluation data, the porcine 50K panel shows high data validity, high data coverage and uniformity of target region and good genotyping effect on DNBSEQ-T7*. Molbreeding and MGI are strategic partners in agricultural molecular breeding. The introduction of high speed, flexible and ultra-high throughput DNBSEQ-T7* and DNBSEQ-400* plaftforms will empowerMolbreeding's need for molecular breeding.
*Unless otherwise stated, all sequencers and sequencing reagents are not available in Germany, USA, Spain, UK, Hong Kong, Sweden, Belgium, and Italy.

 Sequencer Products: SEQ ALL
Sequencer Products: SEQ ALL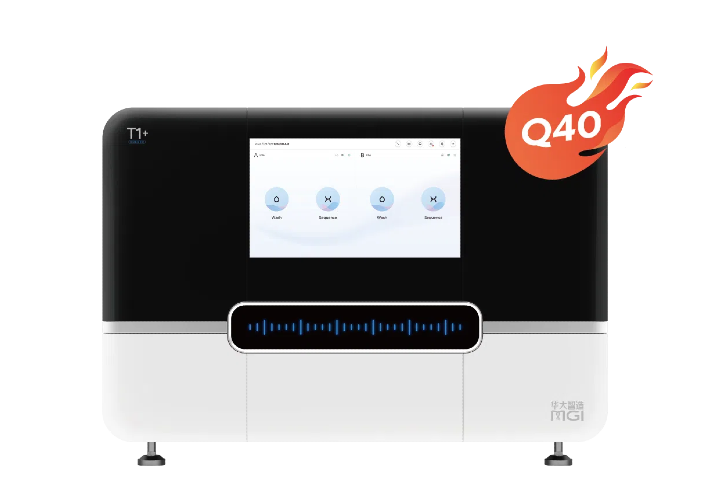











 Technologies
Technologies Applications
Applications Online Resources
Online Resources Data Bulletins
Data Bulletins Service & Support
Service & Support Introduction
Introduction Newsroom
Newsroom Doing Business With Us
Doing Business With Us Creative Club
Creative Club